Published by Bindi M. Doshi, PhD on Nov 16, 2016 9:01:39 AM
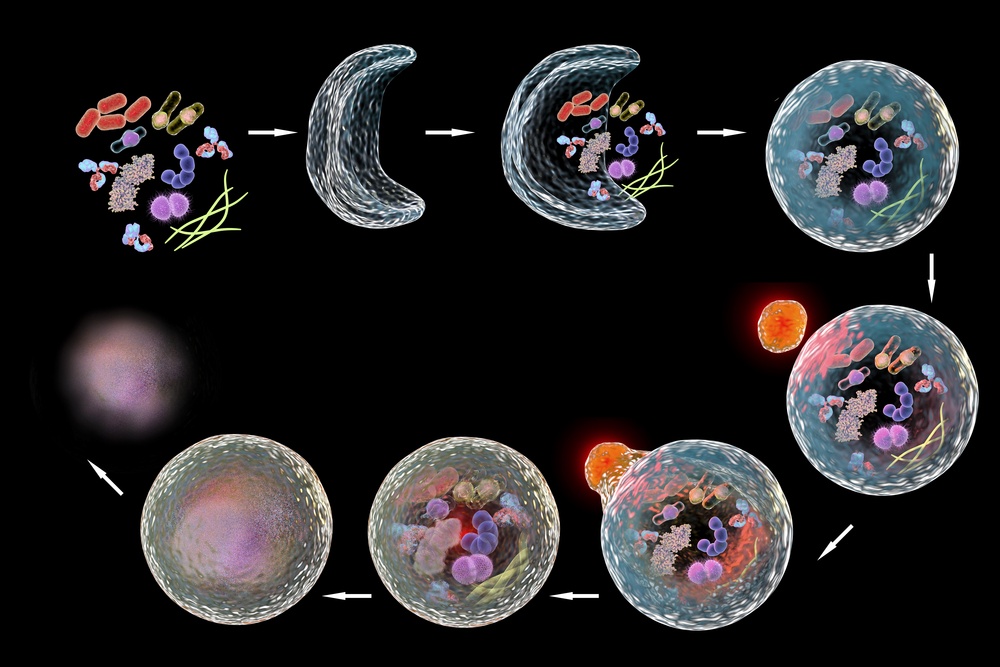
The process of autophagy was first recognized in the late 1950s and it was thought of as bulk “junk” removal. Then, dedicated scientists did a little more digging and found there is an amazing methodology to the “junk” removal. It was discovered that there is rhyme and reason for how a cell decides if and when its components should undergo degradation. Autophagy is rather specific and aids in cell survival by making sure the cell has essential components during times of flux. Autophagy can be selective (i.e. mitophagy) or non-selective (i.e. starvation-induced). Using this method, the cell can have non-essential components recycled or send them to onto autophagolysosomes to be degraded. It is more efficient for a cell to be able to recycle as much as it can, rather than to wait for new proteins to come into the picture. Autophagy is also crucial for clearing out “junk” such as misfolded or aggregated proteins. Research in autophagy has helped provide a better understanding for how a cell is able to survive even under poor conditions.
Early research in yeast helped identify which genes are involved in the autophagy process. These genes are highly conserved in yeast, plants and mammals- a clear indication to the importance of autophagy in all organisms.1 Current research is always evolving and advancing the understanding and implication of autophagy during normal cell function and in disease states.
Autophagy has a few different steps. First there is induction, followed by the initiation phase, where a double membrane starts forming around components to be removed. Elongation and completion seal membrane formation around the material that needs to be degraded, this is now known as autophagosome. Maturation, followed by fusion with lysosome results in the autophagolysosome complex. The final step is degradation.4,5
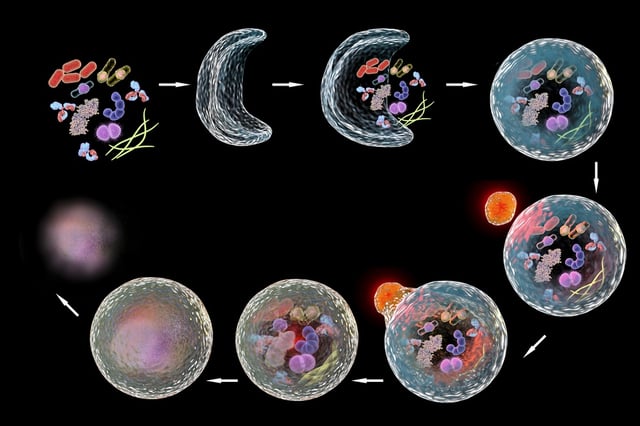
Autophagy is involved in many disease states including cancer, neurodegenerative diseases, and aging. Understanding how selective and induced autophagy are regulated could be a crucial key for potential disease prevention or treatment targets.2,3 The 2016 Nobel Prize in Medicine was awarded for autophagy research. Clearly, the committee recognized the dedication given to this area, which was once considered to be simple bulk garbage disposal. Thanks to the pioneers in this field we have a much better understanding for how the cell processes material to be degraded and how this process works in several different disease states.
MBL has been supporting autophagy research for over twenty years. Our products have been highly cited and are well loved. Click here to see how we can help you in this prize winning field!
1. Ohsumi, Yoshinori. "Historical Landmarks of Autophagy Research." Cell Research 24.1 (2013): 9-23.
2. Li, Shan, Lingfei Wang, Yongzhou Hu, and Rong Sheng. "Autophagy Regulators as Potential Cancer Therapeutic Agents: A Review." CTMC Current Topics in Medicinal Chemistry 15.8 (2015): 720-44.